Assistant Professor, Harvard Medical School
Ph.D., Molecular and Cell Biology, University of California, Berkeley
B.A., Molecular and Cell Biology, University of California, Berkeley
RESEARCH
We study key transcriptional and other gene regulatory events that lead to the acquisition and maintenance of pluripotency in embryonic stem cells (ESCs). ESCs can self-renew or differentiate to produce most of the cells of the body. These distinct but developmentally relevant cell fates are defined by their unique gene expression signatures. Proper execution of these developmental programs requires the precise tuning of gene expression by transcription factors, coactivators and corepressors. Indeed, aberrant transcriptional regulation is the root of many human diseases including developmental disorders, cancers and degenerative diseases.
We have devised various strategies to isolate critical cellular machinery responsible for transcription and to reconstruct the complex process of gene activation in vitro. This technique, in combination with in vivo systems, provides powerful tools to dissect the molecular mechanisms of pluripotency and self-renewal of ESCs, lineage commitment during ESC differentiation, cellular reprogramming, and diseases at the transcriptional level.
Biochemistry of pluripotency and cellular reprogramming
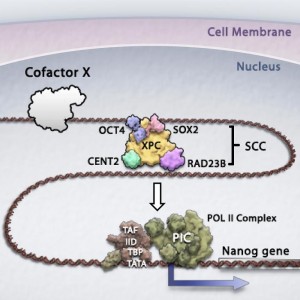
We are in the process of characterizing the in vitro biochemical activities and in vivo functions of three newly identified coactivators essential for transcriptional activation by key stem cell-specific transcription factors OCT4 and SOX2 (Fong et al, Cell, 2011; Fong et al, eLife, 2014). We are also investigating the role of one of these coactivators in the epigenetic regulation of gene expression in pluripotent stem cells, by promoting active DNA demethylation in a genome-wide and locus-specific manner (Ho et al, Genes & Development, 2017). A deeper understanding of the full repertoire of transcription factors operating in stem cells is critical to realizing the full potential of stem cells in tissue regeneration and replacement therapy.
Gene expression and disease mechanisms
Another major focus of the laboratory is to understand how misregulation of transcription can cause human diseases. Using a combination of in vitro system such as directed differentiation of human ESCs, and in vivo model such as zebrafish (in collaboration with Dr. Calum MacRae lab), we have uncovered a previously unrecognized role of a ubiquitously expressed transcription factor in regulating heart function. We aim to identify the critical gene regulatory pathways controlled by this transcription factor, and how misregulation of these gene expression programs leads to tissue-specific disorders such as cardiac arrhythmia and dilated cardiomyopathy.
To contact Dr. Fong, click here.